Modulatory Effect of Jatropha tanjorensis Leaf Extract on the Expression of HOXA11, MECOM, and NFE2 Genes in an Animal Model
| Received 28 Nov, 2025 |
Accepted 01 Apr, 2026 |
Published 07 Apr, 2026 |
Background and Objective: Jatropha tanjorensis leaf extract is traditionally used as medications in the buildup of red blood cells and other cell lines. Homeobox A11(HOXA11), MECOM gene, and NFW2 genes are vital in haemopoiesis. This study is aimed at investigating the expression of these genes and exploring the roles they play in the haemopoietic process. Materials and Methods: A total of thirty (30) male adult albino Wistar rats were selected into 5 groups. The groups were control group (Group 1), rats administered 40 mg phenyl hydrazine (Group 2) to induce anaemia, rats administered with 1000 mg/kg (Group 3), rats administered 3000 and 4000 mg/kg leave extract as group 4 and group 5 respectively. Haematological parameters and MRNA expressions of HOXA 11, MECOM and NFE2 genes were determined using Haematology analyzer and polymerase chain reaction respectively. Data were analyzed using SPSS v21.0 (IBM, USA) using ANOVA, with significance set at p<0.05. Results: The MRNA expression of HOXA 11 was significantly lower in the 1000, 2000 and 4000 mg/kg compared to the control group (p<0.05). MECOM gene expression was significantly higher in the 1000 mg/kg administered group. It increased significantly in the 2000 mg/kg administered group but reduced significantly in the 4000 mg/kg administered group but were all significantly higher compared to the control group (p<0.001). While, there was no significant difference in the 2000 mg/kg administered group compared to the controls (p<0.05). It was significantly lower in the 1000 mg/kg group and further lower in the 4000 mg/kg administered group. Conclusion: Jatropha tanjorensis leaf extract significantly affects the expression of Homeobox A11 (HOXA11), MECOM, and NFE2 genes at varying concentrations of leaf extract and this will significantly impact the haemopoietic process.
| Copyright © 2026 Odaburhine and Syntyche. This is an open-access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. |
INTRODUCTION
The frequency of usage of traditional medicine in developing countries is about 40-90% according to the World Health Organization1. Additionally, many individuals (even in advanced nations) are now turning to plant-based diets and therapeutic options as a result of the increased prevalence of chronic metabolic disorders around the world2. This is connected to bad lifestyle choices and growing urbanization3. This is primarily because many plants and plant-based diets have several benefits that have been recorded, and because they are seen as a safer option than the manufactured supplements and medications4.
Jatropha tanjorensis is a medicinal plant belonging to the Euphorbiaceae family, which is native to certain regions in Africa and Asia. It has been traditionally used in folk medicine for its various therapeutic properties5. Some common names of the plant include: Tanjore Jatropha, Tuba Gbangba (Nigeria), Balubaum (Cameroon), Daafi (Ghana). Jatropha tanjorensis is native to certain regions in Africa and Asia. It is commonly found in tropical and subtropical climates, where it thrives in well-drained soils and receives adequate sunlight6. The plant is known to contain a wide range of biologically active compounds, such as alkaloids, flavonoids, tannins, terpenoids, and saponins, among others3. These chemical constituents have been attributed to its potential medicinal benefits, including antioxidant, anti-inflammatory, antimicrobial, and anticancer activities6. Jatropha tanjorensis is classified as a dicotyledonous plant, which means it belongs to the group of flowering plants that produce two cotyledons (seed leaves) during germination. It is characterized by its woody stem and broad, green leaves2.
The leaves of Jatropha tanjorensis are large, simple, and alternate in arrangement along the stems. They are generally broad and ovate, with a pointed tip and a smooth or slightly toothed margin. The leaf surface is typically dark green, and the veins are prominently visible. The leaves of this plant contain various bioactive compounds responsible for its medicinal properties.
In recent years, there has been growing interest in investigating the pharmacological properties of Jatropha tanjorensis and its potential use in modern medicine3. Several studies have explored its effects on various physiological processes, such as wound healing, immune modulation, and pain relief7. However, there is limited scientific literature on its specific impact on gene expression, especially in the context of important regulatory genes like HOXA11, MECOM, and NFE28.
Haematopoiesis, also known as haematopoiesis, is the process by which blood cells are formed and differentiated from hematopoietic stem cells (HSCs) in the bone marrow9. This highly regulated and complex process is essential for maintaining a constant supply of functional blood cells that are necessary for oxygen transport, immune response, and haemostasis. The expression and activity of various genes play crucial roles in regulating haematopoiesis at different stages. This study will examine the effect of three specific genes, HOXA11, MECOM, and NFE2, on haematopoiesis in detail.
HOXA11 is a member of the homeobox gene family, which encodes transcription factors that are involved in the regulation of developmental processes. During haematopoiesis, HOXA11 has been found to play critical roles in the formation and patterning of blood cell lineages. Its expression is dynamically regulated during different stages of haematopoiesis.
Studies have shown that HOXA11 is involved in the differentiation of erythroid cells, which are responsible for carrying oxygen in the blood9. It contributes to the formation of definitive erythroid cells by promoting their maturation and controlling the switch from primitive to definitive erythropoiesis during embryonic development10. Additionally, HOXA11 is implicated in regulating megakaryocyte differentiation and platelet production, influencing thrombopoiesis and haemostasis10. The disruption or dysregulation of HOXA11 expression can lead to abnormalities in blood cell development, potentially causing haematological disorders such as anaemia, thrombocytopenia, or platelet dysfunction9.
MECOM (EVI1) also known as EVI1 (Ecotropic Viral Integration Site 1), is a transcription factor that plays a complex and dual role in haematopoiesis9. Depending on the cellular context and stage of differentiation, MECOM can either promote or inhibit haematopoiesis11. During early embryonic development, MECOM is essential for maintaining the self-renewal capacity of hematopoietic stem cells12. It prevents their differentiation, thereby ensuring a steady pool of undifferentiated stem cells. However, as haematopoiesis progresses and stem cells commit to differentiate into specific blood cell lineages, MECOM expression needs to be downregulated to allow for proper lineage commitment and differentiation13.
MECOM has been implicated in various haematological malignancies. In some cases, it is overexpressed, leading to the inhibition of normal haematopoiesis and promoting uncontrolled proliferation of leukemic cells. On the other hand, in certain leukaemias, MECOM’s expression might be silenced or disrupted, contributing to the loss of cell differentiation control and abnormal expansion of immature blasts cells14.
NFE2 (Nuclear Factor, Erythroid 2) is a transcription factor that is particularly crucial for the maturation of erythroid cells13. It regulates the expression of antioxidant genes and plays a vital role in protecting erythroid cells from oxidative stress, which is essential for their survival and functionality during the oxygen-carrying process13. During erythropoiesis, NFE2 regulates the expression of genes involved in heme biosynthesis, globin chain synthesis, and iron metabolism, all of which are essential for the proper maturation and function of red blood cells. NFE2 also participates in the differentiation of other blood cell lineages, such as megakaryocytes and platelets.
Mutations or disruptions in NFE2 have been associated with various haematological disorders, including anaemia and myeloproliferative disorders. Dysregulation of NFE2 can lead to impaired erythropoiesis, resulting in anaemia or ineffective erythropoiesis15.
These genes: HOXA11, MECOM (EVI1), and NFE2 play crucial roles in regulating different aspects of haematopoiesis, from stem cell maintenance and lineage commitment to the differentiation and maturation of specific blood cell types11. Dysregulation of these genes can have significant implications for normal haematopoiesis and may contribute to the development of various haematological disorders. Therefore, research into the molecular mechanisms controlled by these genes will provide valuable insights into their potential therapeutic applications in the context of haematological diseases13.
This study aims to investigate the modulatory effect of Jatropha tanjorensis leaf extract on the expression of HOXA11, MECOM, and NFE2 genes in an animal model.
MATERIALS AND METHODS
Study area: The study was carried out meticulously at the University of Benin, Benin City, Edo State, Nigeria. The study duration for this work was March-August, 2023.
Identification of the Jatropha tanjorensis: Jatropha tanjorensis leaves were harvested from the University of Benin environs, Benin City, Edo State, Nigeria. It was further transported to the Department of Plant Biology and Biotechnology, Faculty of Life Sciences, University of Benin, and it was given a voucher number UBH-G532.
Animal care: The animals were housed in a cross-ventilated room in the animal holdings of the Department of Anatomy, University of Benin, Benin City. Animals were exposed to 12 hrs of dark and light cycles with access to feed and water ad libitum. The rats were acclimatized for a period of 2 weeks and fed using a pre-determined concentration of Jatropha tanjorensis. Ethical approval was obtained from the Edo State Ministry of Health, Nigeria, with approval No. HA/737/23/D/08290158.
Study design: Thirty male adult albino Wistar rats were selected into group 1 (control) and three experimental groups, 2, 3, and 4. Each group consists of 6 animals. Afterwards, they were fed for 1 month following the established feed formulation.
| • | Group 1: This was the control group. Animals in this group received only standardized feed and water ad libitum | |
| • | Group 2: Animals in this group were administered 1000 mg/kg of Jatropha tanjorensis crude extract | |
| • | Group 3: Animals in this group were administered 2000 mg/kg of Jatropha tanjorensis crude extract | |
| • | Group 4: Animals in this group were administered 4000 mg/kg of Jatropha tanjorensis crude extract. |
The experimental period lasted for 5 weeks
Sacrifice of the animals and sample collection: At the end of the experimental period, the animals were grossly observed for general physical characteristics and were weighed using a weighing balance. A midline incision was made through the ventral wall of the rats after anaesthetizing using chloroform. Bone marrow samples were collected from the femur of the rats and stored in 0.3 mL of TriZol reagent14.
Laboratory analysis
Phytochemical analysis (crude extraction): Three different solvents (n-hexane, methanol, and water) were used for the extraction of Jatropha tanjorensis based on their polarity. Three hundred 300 g of the pulverized leaf of Jatropha tanjorensis was placed in three 1000 mL beakers. The solvents were then added and allowed to soak for 3 days. After 3 days, the extract was filtered and the solution was placed in a freeze-dryer so that a dry paste of the compound could be obtained. The freeze-drying process aimed to remove water content from the extract, yielding a dry paste or solid residue. The freeze dryer operates by subjecting the solution to low temperatures under vacuum conditions, causing the frozen water to sublimate directly from ice to vapour.
High-performance liquid chromatography (HPLC) analysis: High-Performance Liquid Chromatography (HPLC) is a method used to separate a mixture of complex samples. HPLC is an active process in which materials are pumped at high pressure through a separation column, which contains a stationary phase, usually a chemically functional bead that separates the compound mixtures. Samples are introduced through the injector and carried with the mobile phase across the stationary phase to affect the separations. After separation through the column, the samples were exposed to a detector system that identifies and quantifies the individual compounds.
Extraction: In the experimental procedure, a precise amount of 1 g of the sample was meticulously weighed and placed in an amber bottle. Subsequently, a mixture comprising 20 mL of acetonitrile and methanol was introduced to the sample within the bottle, and the entire concoction was vigorously shaken for a duration of 30 min. Following the vigorous shaking process, the aqueous portion was allowed to naturally separate and drain, while the organic solvent phase was collected into a standard flask with a capacity of 25 mL. This collected organic solvent was meticulously brought up to the specified level on the flask, rendering it ready for subsequent analysis. To establish a baseline and reference for the analytes being studied, a standardized profile of the analytes was initially injected into the High-Performance Liquid Chromatography (HPLC) system. This injection generated a chromatogram that exhibited distinct peaks with corresponding peak areas and profiles. These parameters were then employed to configure the HPLC system for the forthcoming analysis of the test samples.
Subsequently, a carefully measured portion of the extracted test sample was introduced into the HPLC system for analysis. This injection resulted in the generation of a chromatogram that displayed a distinctive peak area and profile, which corresponded to the analytes present in the sample. By comparing the peak area of the sample with that of the previously established standard, and considering the concentration of the standard, the concentration of the test sample was accurately determined.
RNA extraction and semi-quantitative polymerase chain reaction (PCR): The RNA was isolated from the rat bone marrow with TRIzol Reagent (ThermoFisher Scientific) and converted to cDNA using the ProtoScript First Strand cDNA Synthesis Kit (NEB). PCR amplification of HOXA11 mRNA, MECOM mRNA, and NFE2 mRNA was done using OneTaq® 2X Master Mix (NEB).
After sacrificing the animals, the bone marrow harvested was embedded in 0.3 mL of TRIzol (Invitrogen Life Technologies, Inc., Carlsbad, CA) reagent inside an Ependorf tube for proper tissue preservation. The tissues were homogenized using a plastic pestle. The RNA lyase buffer was then added to the homogenate to further break down the tissue cell membranes, after which it was spun at 10000 rpm for 10 min. The supernatant, which contains the RNA, was carefully removed and placed in a separate Eppendorf tube. The RNA precipitating buffer was then centrifuged at 10000 rpm for 30 min. The supernatant was carefully removed remaining the RNA precipitate at the bottom of the tube. The RNA wash buffer will be added and centrifuged again at 10000 rpm for 5 min. This step was repeated 3 times to remove excess solutions and buffers previously added. Nuclease-free water was added to break down the RNA into a pellet form and the phosphodiester bond of the RNA. The solution also has a nuclease inhibitor, which removes other DNA or RNA contaminants from the medium. The RNA was then quantified using a UV spectrophotometer at 260 nm.
Complementary DNA (cDNA) synthesis: All components of the cDNA kit were added to the RNA following the manufacturer’s instructions; these components included random primer, oligonucleotide, primer or deoxynucleotides, and reverse transcriptase buffer. After adding all components to the RNA, the mixture was then incubated at 420°C in a thermocycler for 1 hour. Then the temperature was increased to 750°C to denature the reverse transcriptase. Thereafter, all the RNA was converted to cDNA. For RQ-PCR on the rat sample, the expression levels of all six genes were normalized to the levels of the GADPH housekeeping gene.
Gene amplification: An equal volume of both forward and reverse primers was added, PCR mix (master mix), Taq polymerase, and magnesium was also added. The mixture was placed in a thermo-cycler for amplification. The thermo-cycler is programmed for 30 cycles.
Gel electrophoresis: After the PCR process, the DNA gel loading dye was added to the mixture. The agarose gel was prepared by dissolving 1% of agarose gel in TBE buffer. The gel was then allowed to solidify, and then the sample was loaded, and the gel was connected for electrophoresis. Thereafter, a snapshot was taken. The image was then transferred to ImageJ. The quantification of band intensity was done using “ImageJ” software.
Statistical analysis: Data obtained from this research were presented and analysed using statistical package for the Social Sciences (SPSS) version 21.0 (IBM Inc., USA). Analysis of variance (ANOVA) was used to compare treatment groups of continuous variables. Bar chats was used to represent the mRNA gene expression patterns. A p<0.05 was considered statistically significant.
RESULTS
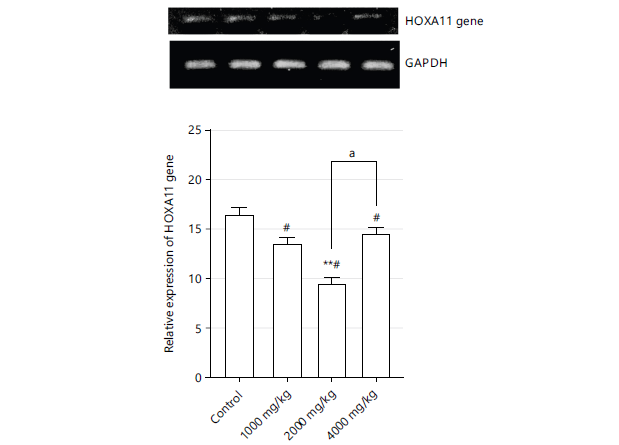
Figure 1 shows the expression of genes as represented by gel electrophoresis photomicrograph of control, 1000, 2000, and 4000 mg/kg groups. The mRNA expression of HOXA11 is significantly lower (p>0.001) in the 2000 mg/kg group compared to the control group. HOXA11 gene expression in the 4000 mg/kg group was significantly higher (p<0.05) compared to the 2000 mg/kg group but significantly lower (p<0.05) compared to the untreated group.
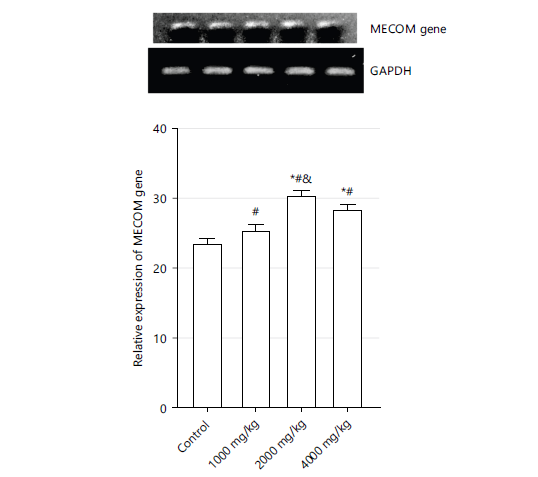
Figure 2 shows the expression of the MECOM gene as represented by gel electrophoresis at 1000, 2000, and 4000 mg/kg. The expression of the MECOM gene in 1000, 2000, and 4000 mg/kg were all significantly higher (p<0.05) when compared to the control group. Both 2000 and 4000 mg/kg showed higher significance compared to the control group.
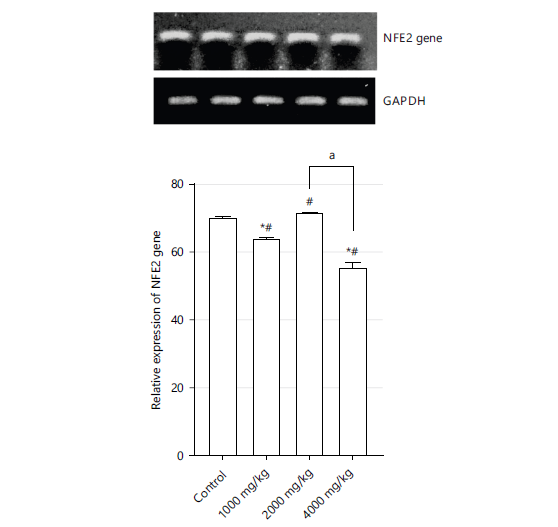
Figure 3 shows the expression of the NFE2 gene as represented by gel electrophoresis at 1000, 2000, and 4000 mg/kg. Results show significantly lower (p<0.05) expressions of the NFE2 gene in 1000 and 4000 mg/kg compared to the control group. A 1000 and 2000 mg/kg show significantly higher expressions compared to the control group. The 4000 mg/kg group shows lower significant expressions compared to the 2000 mg/kg group.
|
|
|
DISCUSSION
Jatropha tanjorensis, a plant belonging to the Euphorbiaceae family, has garnered attention for it’s potential bioactive compounds with diverse pharmacological properties. Extracts from various parts of the plant, including leaves, stems, and roots, have been investigated for their medicinal potential15. This study was carried out to assess the modulatory effect of this leaf extract in the expression of Homeobox A11 (HOXA11), MECOM, and NFE2 genes using albino Wistar rats.
The HOX A11 genes a protein-coding gene located on human chromosome 7, plays a crucial role in embryonic development, and was significantly lower in the group treated with 1000 mg/kg leave extract of Jatropha tanjorensis compared to the controls. This indicates that the leaf extract may not have a direct effect on the function of these genes. Although some studies have shown the presence of a range of metabolites such as alkaloids, flavonoids, terpenoids, and phenolic compounds, which are believed to contribute to its biological activities16 these may not have any effect on the expression of this gene. The HOXA11 gene was further lowered when the concentration of leave extract was increased to 2000 mg/kg. This may indicate a downregulation and inhibitory effect at a higher concentration. On administration of the 4000 mg/kg concentration, the expression of this gene was significantly higher (p<0.05). This study did not show a steady increase or a decrease when varying concentrations of the leaf extract were administered, but rather it showed varying changes in the different doses of leaf extract administered. These varying changes in the result observed may be attributed to physiological changes in the rats and also the adaptive and environmental response of the rats to the different doses of the leaf extract16.
The MECOM gene, which plays a crucial role in various biological processes including blood cell development (Haematopoiesis), cell proliferation and differentiation, was observed to be significantly higher (p<0.05) in the group administered with 1000 mg/kg of leaf extract when compared with the controls. This indicates that the leaf extract is able to stimulate the activity of this gene. On administration of 4000 mg/kg the expression of MECOM gene was significantly higher but it reduced with the administration of 4000 mg/kg. Generally, the administration of the leave extract significantly increased the expression of the MECOM genes when compared to the control groups. This indicates the activation of this gene by the varying concentrations of Jatropha tanjorensis leaf extract.
Nuclear Factor Erythroid 2 (NFE2) gene, a transcription factor that is particularly responsible for the maturation of erythroid cells13 was significantly lower with the administration of 1000 mg/kg leave extract, but was significantly higher with the administration of 2000 mg/kg leaf extract of Jatropha tanjorensis. This may be attributed to the fact expression of NFE2 genes requires a moderate administration of the leaf extract. It also signifies that too low a concentration and too high a concentration of leaf extract are inhibitory to the expression of this gene, as shown in Fig. 3. This further implies that moderate administration of leave extract can increase the expression of this gene, which will invariably encourage proliferation of erythroid cells and help in erythropoiesis. During erythropoiesis, NFE2 regulates the expression of genes involved in heme biosynthesis, globin chain synthesis, and iron metabolism, all of which are essential for the proper maturation and function of red blood cells. The NFE2 also participates in the differentiation of other blood cell lineages, such as megakaryocytes and platelets.
CONCLUSION
The findings of this study indicate that Jatropha tanjoresis leaf extract has an impact on the expression of some genes that help in haematopoiesis, thereby leading to the formation of various blood cell lines. It demonstrates that different doses of leave extract administration affect the expression of HOXA11, MECOM, and NFE2 genes. Moderate doses of 2000 mg/kg leave extract lead to significantly higher expression of HOXA11, MECOM, and NFE2 genes, but lower and higher doses of leave extracts are inhibitory to the expression of these genes. It is concluded that moderate administration of Jatropha tanjorensis leaf extract will encourage erythropoiesis as observed in this study.
SIGNIFICANCE STATEMENT
This study provides molecular evidence supporting the haemopoietic relevance of Jatropha tanjorensis by demonstrating its dose-dependent modulation of key regulatory genes (HOXA11, MECOM, and NFE2). The findings clarify possible genetic pathways through which the leaf extract influences blood cell formation, thereby strengthening its traditional therapeutic claims and offering a scientific basis for its potential development as a supportive agent in anaemia management and haemopoietic research.
REFERENCES
- Wang, M., L. Zheng, R. Lin, S. Ma, J. Li and S. Yang, 2023. A comprehensive overview of exosome lncRNAs: Emerging biomarkers and potential therapeutics in endometriosis. Front. Endocrinol., 14.
- Bury, L., E. Falcinelli and P. Gresele, 2021. Learning the ropes of platelet count regulation: Inherited thrombocytopenias. J. Clin. Med., 10.
- Stephenson, K., L. Kennedy, L. Hargrove, J. Demieville, J. Thomson, G. Alpini and H. Francis, 2017. Updates on dietary models of nonalcoholic fatty liver disease: Current studies and insights. Gene Expression, 18: 5-17.
- Kim, C.S., S. Park and J. Kim, 2017. The role of glycation in the pathogenesis of aging and its prevention through herbal products and physical exercise. J. Exercise Nutr. Biochem., 21: 55-61.
- Chibuogwu, C.C., U.O. Njoku, F.C.O. Nwodo, E.O.V. Ozougwu and N.V. Nweze, 2021. Toxicity assessment of the methanol extract of Jatropha tanjorensis (Euphorbiaceae) leaves. Future J. Pharm. Sci., 7.
- Oladele, J.O., O.T. Oladele, A.O. Ademiluyi, O.M. Oyeleke, O.O. Awosanya and O.I. Oyewole, 2020. Chaya (Jatropha tanjorensis) leafs protect against sodium benzoate mediated renal dysfunction and hepatic damage in rats. Clin. Phytosci., 6.
- Danborno, A.M., F. Tarfa, J.E. Toryila, E.U. Awheela and V.T. Shekarau, 2019. The effects of Jatropha tanjorensis aqueous leaf extract on haematological parameters in Wistar rats. J. Afr. Assoc. Physiol. Sci., 7: 133-137.
- Cuellar-Nuñez, M.L., I. Luzardo-Ocampo, R. Campos-Vega, M.A. Gallegos-Corona, E.G. de Mejía and G. Loarca-Piña, 2018. Physicochemical and nutraceutical properties of moringa (Moringa oleifera) leaves and their effects in an in vivo AOM/DSS-induced colorectal carcinogenesis model. Food Res. Int., 105: 159-168.
- Binns, T.C., C.A. Tormey and H.M. Rinder, 2022. Platelet Production and Kinetics. In: Rossi's Principles of Transfusion Medicine, Simon, T.L., E.A. Gehrie, J. McCullough, J.D. Roback and E.L. Snyder, Wiley, New Jersey, ISBN: 9781119719809, pp: 158-167.
- Jin, S., J. Li, M. Barati, S. Rane and Q. Lin et al., 2020. Loss of NF-E2 expression contributes to the induction of profibrotic signaling in diabetic kidneys. Life Sci., 254.
- Tang, M., X. Bai, J. Wang, T. Chen and X. Meng et al., 2021. Efficiency of graft-transmitted JcFT for floral induction in woody perennial species of the Jatropha genus depends on transport distance. Tree Physiol., 42: 189-201.
- di Zhang, S. Iwabuchi, T. Baba, S.I. Hashimoto, N. Mukaida and S.I. Sasaki, 2020. Involvement of a transcription factor, Nfe2, in breast cancer metastasis to bone. Cancers, 12.
- Wang, D., S. Wang, Z. Zhou, D. Bai and Q. Zhang et al., 2022. White blood cell membrane-coated nanoparticles: Recent development and medical applications. Adv. Healthcare Mater., 11.
- Bae, J., T. Hideshima, G.L. Zhang, J. Zhou, D.B. Keskin, N.C. Munshi and K.C. Anderson, 2018. Identification and characterization of HLA-A24-specific XBP1, CD138 (Syndecan-1) and CS1 (SLAMF7) peptides inducing antigens-specific memory cytotoxic T lymphocytes targeting multiple myeloma. Leukemia, 32: 752-764.
- Falodun, A., A.A. Udu-Cosi, O. Erharuyi, V. Imieje, J.E. Falodun, O. Agbonlahor and M.T. Hamann, 2013. Jatropha tanjorensis-review of phytochemistry, pharmacology, and pharmacotherapy. J. Pharm. Allied Sci., 10: 1955-1964.
- Kaluzhny, Y., K. Ravid and M. Poncz, 2003. Transcription Factors Involved in Lineage‐specific Gene Expression During Megakaryopoiesis. In: Transcription Factors: Normal and Malignant Development of Blood Cells, Ravid, K. and J.D. Licht (Eds.), Wiley, New Jersey, ISBN: 9780471223887, pp: 31-49.
How to Cite this paper?
APA-7 Style
Odaburhine,
O.E., Syntyche,
O.O. (2026). Modulatory Effect of Jatropha tanjorensis Leaf Extract on the Expression of HOXA11, MECOM, and NFE2 Genes in an Animal Model. Science International, 14(1), 41-49. https://doi.org/10.17311/sciintl.2026.41.49
ACS Style
Odaburhine,
O.E.; Syntyche,
O.O. Modulatory Effect of Jatropha tanjorensis Leaf Extract on the Expression of HOXA11, MECOM, and NFE2 Genes in an Animal Model. Sci. Int 2026, 14, 41-49. https://doi.org/10.17311/sciintl.2026.41.49
AMA Style
Odaburhine
OE, Syntyche
OO. Modulatory Effect of Jatropha tanjorensis Leaf Extract on the Expression of HOXA11, MECOM, and NFE2 Genes in an Animal Model. Science International. 2026; 14(1): 41-49. https://doi.org/10.17311/sciintl.2026.41.49
Chicago/Turabian Style
Odaburhine, Osime, Evarista, and Omoruyi Osamiomwanfan Syntyche.
2026. "Modulatory Effect of Jatropha tanjorensis Leaf Extract on the Expression of HOXA11, MECOM, and NFE2 Genes in an Animal Model" Science International 14, no. 1: 41-49. https://doi.org/10.17311/sciintl.2026.41.49

This work is licensed under a Creative Commons Attribution 4.0 International License.


